|
|
|
|
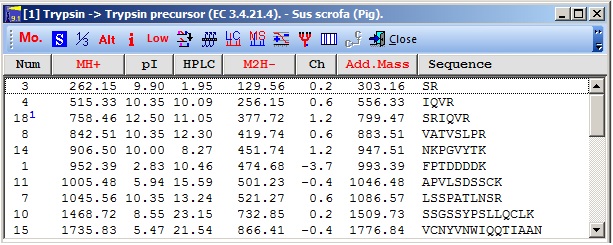
Peptide window |
|||||
 |
|||||
|
The peptide window lists all the peptides that will be generated from the given protein along with a large number (>20) of physical-chemical parameters Peptides can be generated with partial cleavages (indicated by the blue superscript in the picture), modified termini, modified residues (even partial modifications are supported in a limited way). Specific residues that are colored for easier referencing in the sequence window is carried through to this window.. Sorting can take place on any column by clicking on the header. Clicking a second time reverses the sort. Through the toolbar and/or the pop-up menu (right mouse click) you can access additional functions related either to the digest (like the simulated HPLC reversed phase chromatogram) or to the currently selected peptide (ms/ms cleavage, peptide information, charge vs. pH graph). |
|||||
|
Site last updated: February 14, 2025 |
|||||
